| p-value: | 1e-8 |
| log p-value: | -1.896e+01 |
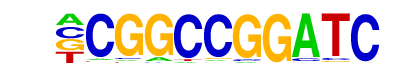
| Information Content per bp: | 1.809 |
| Number of Target Sequences with motif | 39.0 |
| Percentage of Target Sequences with motif | 17.11% |
| Number of Background Sequences with motif | 2793.4 |
| Percentage of Background Sequences with motif | 6.10% |
| Average Position of motif in Targets | 1289.1 +/- 487.1bp |
| Average Position of motif in Background | 1051.4 +/- 673.0bp |
| Strand Bias (log2 ratio + to - strand density) | 0.0 |
| Multiplicity (# of sites on avg that occur together) | 1.16 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
Elk4(ETS)/Hela-Elk4-ChIP-Seq(GSE31477)/Homer
| Match Rank: | 1 |
| Score: | 0.61
| | Offset: | 2
| | Orientation: | reverse strand |
| Alignment: | CGGCCGGATC--
--RCCGGAARYN |
|


|
|
SPDEF(ETS)/VCaP-SPDEF-ChIP-Seq(SRA014231)/Homer
| Match Rank: | 2 |
| Score: | 0.60
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | CGGCCGGATC-
-ANCAGGATGT |
|


|
|
POL013.1_MED-1/Jaspar
| Match Rank: | 3 |
| Score: | 0.60
| | Offset: | 4
| | Orientation: | reverse strand |
| Alignment: | CGGCCGGATC
----CGGAGC |
|

|
|
POL011.1_XCPE1/Jaspar
| Match Rank: | 4 |
| Score: | 0.59
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | CGGCCGGATC
GGGCGGGACC |
|

|
|
Fli1(ETS)/CD8-FLI-ChIP-Seq(GSE20898)/Homer
| Match Rank: | 5 |
| Score: | 0.58
| | Offset: | 2
| | Orientation: | reverse strand |
| Alignment: | CGGCCGGATC--
--DCCGGAARYN |
|


|
|
PB0077.1_Spdef_1/Jaspar
| Match Rank: | 6 |
| Score: | 0.58
| | Offset: | -3
| | Orientation: | forward strand |
| Alignment: | ---CGGCCGGATC---
GTACATCCGGATTTTT |
|

|
|
Elk1(ETS)/Hela-Elk1-ChIP-Seq(GSE31477)/Homer
| Match Rank: | 7 |
| Score: | 0.58
| | Offset: | 2
| | Orientation: | reverse strand |
| Alignment: | CGGCCGGATC--
--RCCGGAAGTD |
|


|
|
E2F6(E2F)/Hela-E2F6-ChIP-Seq(GSE31477)/Homer
| Match Rank: | 8 |
| Score: | 0.58
| | Offset: | 1
| | Orientation: | forward strand |
| Alignment: | CGGCCGGATC-
-GGCGGGAARN |
|


|
|
E2F1(E2F)/Hela-E2F1-ChIP-Seq(GSE22478)/Homer
| Match Rank: | 9 |
| Score: | 0.56
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -CGGCCGGATC
CWGGCGGGAA- |
|

|
|
MA0470.1_E2F4/Jaspar
| Match Rank: | 10 |
| Score: | 0.55
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | CGGCCGGATC-
GGGCGGGAAGG |
|


|
|